Running the experiments¶
If you have set up the databases and the external software, you are ready to run the face recognition experiments. Don’t worry if you couldn’t download all the databases, at least some experiments can be executed. Of course, you can only run those experiments, for which you have the images.
To execute an experiment, there are several scripts in the bin directory, which are explained in more detail in this section. All experiments are based on the scripts defined in bob.bio.base, which executes a sequence of processes. The scripts in this package only provide the command line options for ./bin/verify.py and ./bin/grid_search.py.
Warning
Some of the experiments require a long execution time (several days or weeks) and several Gigabyte of memory (up to approx. 80 GB).
Each face recognition experiment is sub-divided into three steps:
- Pre-processing: geometrical alignment and photometric enhancement of the source images.
- Face recognition: feature extraction, model enrollment and scoring.
- Evaluation: interpreting the scores and generating plots.
Usually, image pre-processing and face recognition build an experiment in bob.bio, but since the preprocessed images are shared between several algorithms, in this package we split off the pre-processing as an extra step.
Common interface¶
All scripts that are discussed in this section have a common set of command line options. Each option has a shortcut (with one dash) and a long version (with two dashes), both of which are identical.
- -h, --help: Lists the command line options, their possible parameters and defaults.
- -x, --task: One of the three steps: preprocess, execute or evaluate. This option needs to be specified.
- -a, --algorithms: a list of algorithm names that should be executed. The actual algorithms might change according to the experiment. By default, all desired algorithms are executed.
- -R, --result-directory: The directory, where resulting score files should be written to. [1]
- -T, --temp-directory: The directory, where temporary files files should be written to. This directory might be relative or absolute. [2]
- -w, --plot: The file that the plot should be written to during evaluation. [3]
- -g, --grid: This option is mainly to be used at Idiap. It will submit all experiments to the SGE grid, instead of running them sequentially on the local machine (which happens when the --grid option is not used). If your institution provides an SGE grid as well, please modify the GridTK package towards your grid to be able to use this option.
- -p, --parallel: This option can be used to run execute the experiments in (up to) the given number of parallel jobs on the local machine. [4]
- -v, --verbose: Increases the verbosity level. It might be specified several times to set the level to (0: ERRORS (the default), 1: WARNINGS, 2: INFO, 3: DEBUG). Usually it is a good idea to have at least INFO level (e.g. by -vv).
- -q, --dry-run: Just write the calls to the the raw script to console, instead of executing them.
In some cases, you might want to pass additional parameters to the raw script directly. To do so, please separate the parameters to the script and the parameters to the raw script with a double dash (--). For a list of parameters that might be passed to the raw script, please refer to its documentation.
Several experiments are run on many databases. In these cases, they have another command line option:
- -d, --database or --databases: Select one or several databases to run the experiments on. See the --help options for valid databases and the default(s).
Some experiments can be evaluated also on the score files of COTS algorithm, which can be downloaded from our biometric resources web page. As an easy way to include the results of the COTS algorithm into the plots is to use the --all flag.
Note
In any of the experiments below, the evaluation procedure can be run even if some of the experiments haven’t finished yet. Any result that is not yet available will be left empty. Hence, you can watch the plots grow when running the evaluation at different times during the experiment.
| [1] | The --result-directory might be specified absolute or relative to the current working directory. Inside this directory, score files will be written by the execute task, and read and evaluated by the evaluate task. If you don’t want to re-run the according experiments, score files are present also from our data website. |
| [2] | The --temp-directory might be specified absolute or relative to the current working directory. This directory needs to have a lot of free space as a lot of temporary data will be written to these directories. Any experiment is designed to check if temporary files are present, and regenerate them, if not. After experiments finished, the temp directory might be deleted. |
| [3] | In most cases this file is a multi-page .pdf file containing several plots. |
| [4] | The parallel option will parallelize each single face recognition experiment, whenever possible, using the specified number of parallel processes on the local machine. Under the hood, it uses the --local option of GridTK. Please be aware that no memory checks will be performed, so if each parallel job requires memory, all processes together might require more memory than your local machine has available. |
Image resolution experiments¶
The first set of experiments executes the all algorithms on images with different resolutions. In these tests, non-optimized configurations of all algorithms are used to determine the image resolution that is best suited for the experiment.
Under the hood, this experiment uses the ./bin/grid_search.py to execute the algorithms. The according configuration file can be found in this package under bob/chapter/FRICE/configurations/image_resolution.py, where the preprocessor and a list of tested image heights is defined. The definitions of the extractors and the algorithms is given in the script bob/chapter/FRICE/script/image_resolution.py, where resolution-specific parameters are modified using the #F parameter.
To execute the image preprocessing algorithm tests, simply call:
$ ./bin/image_resolution -vv -x preprocess
$ ./bin/image_resolution -vv -x execute
$ ./bin/image_resolution -vv -x evaluate
Of course, you need to wait between executing the three commands. Finally, the resulting plot, which corresponds to figure 5 a) of the book chapter, is generated as the file resolutions.pdf.
Note
As indicated above, you can add any other command line parameter. A useful parameter might be --parallel to speed up processing (the parallel option is ignored in the third command, i.e., the evaluation). Also, the --dry-run option might come in handy, i.e., to see the full command lines of the commands, which are actually executed.
Image pre-processing algorithms¶
Secondly, different preprocessing algorithms are tested for each of the face recognition algorithms. For this test, we execute the algorithms on four different databases: banca (protocol P), arface (illumination protocol), multipie (illumination protocol) and xm2vts (darkened protocol) As all other following tests, the image resolution of 64x80 pixels is used.
Under the hood, this experiment also uses the ./bin/grid_search.py to execute the algorithms. The according configuration file can be found in this package under bob/chapter/FRICE/configurations/image_preprocessor.py, where a list of preprocessors is defined. The definitions of the extractors and the algorithms is given in the script bob/chapter/FRICE/script/image_preprocessor.py. This time, the extractors and algorithms do not need to be adapted to the preprocessor.
To execute the image preprocessing algorithm tests, simply call:
$ ./bin/image_preprocessor -vv -x preprocess
$ ./bin/image_preprocessor -vv -x execute
$ ./bin/image_preprocessor -vv -x evaluate
The resulting plot, which corresponds to figures 5 b)-d) of the book chapter, is generated as the multi-page PDF file preprocessor.pdf. An additional plot at the beginning of the PDF shows the average performance of each of the preprocessing algorithms.
Note
By default, the experiments are executed on all 4 databases. However, you might select some of the databases using the --databases command line option (which you have to specify for all three commands). The evaluation will remove the databases from the plot accordingly.
Configuration optimization¶
Now that we have a proper image resolution and an image preprocessing algorithm for each of the tested face recognition algorithms, we want to optimize the configurations of all algorithms. For each algorithm, parameter optimization is done in several steps, where the latter steps always use the optimal parameters obtained in the previous steps. The step can be defined using the --step (or short -t) command line option, and valid values are 1, 2 and 3
Under the hood, once more ./bin/grid_search.py is used to execute the algorithms. Here is a short listing of the algorithms and their optimization steps – and the optimal results that you should obtain.:
- LGBPHS (defined in bob/chapter/FRICE/configuration/LGBPHS.py):
- Size and overlap of image blocks. Best values: 4x4 pixels with no overlap.
- Parameters of the Gabor wavelet transform. Best values:
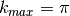 ,
, 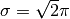 , and Gabor phases included.
, and Gabor phases included. - LBP types and histogram intersection measures. Best values: non-uniform
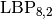 codes compared with histogram intersection.
codes compared with histogram intersection.
- Graphs (defined in bob/chapter/FRICE/configuration/Graphs.py):
- Gabor wavelet parameters and similarity function. Best values:
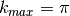 ,
, 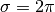 and the
and the 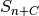 similarity function combining absolute and phase values of Gabor jets.
similarity function combining absolute and phase values of Gabor jets. - Distance between two nodes and handling of several graphs in one model. Best values: 1x1 pixels. [5] and sum over all node positions and take the maximum similarity over all model jets at this node.
- Gabor wavelet parameters and similarity function. Best values:
- ISV (defined in bob/chapter/FRICE/configuration/ISV.py):
- Size and overlap of image blocks. Best values: 10x10 pixels with maximum overlap.
- Number of DCT coefficients. Best value: 45.
- Number of Gaussians and ISV subspace dimension. Best values: 768 and 200. [6]
To run the experiments, simply call:
$ ./bin/configuration_optimization -vv -x preprocess
$ ./bin/configuration_optimization -vv -x execute -t 1
$ ./bin/configuration_optimization -vv -x execute -t 2
$ ./bin/configuration_optimization -vv -x execute -t 3
$ ./bin/configuration_optimization -vv -x evaluate --all
At the end, the evaluation script will generate a list of plots and store them in configuration_optimization.pdf, which show the results of the configuration optimization steps graphically. Unfortunately, there was not enough space in the book chapter to actually display the steps there, which is really sad since it took some time to create the plots and I find them really beautiful...
Note
When the timing experiments have finished, an additional plot will be created during the evaluate task, which shows the results of the optimized algorithms on banca development and evaluation sets. Additionally, the results from LDA-IR and COTS are reported in that plot, which is appended to the end of configuration_optimization.pdf.
| [5] | The results of different node distances were quite similar. Since node distance 1x1 would have generated huge graphs, we decided to go on with the second best option, which was 6x6. |
| [6] | Probably more Gaussians could generate better results, but this would require more memory and longer execution time. We tried to use 1024 Gaussians, but the ISV training exceeded our memory capacity. |
Partial occlusions¶
Now, the configurations for each of the algorithms are optimized. They are stored in bob/chapter/FRICE/configurations/optimized.py, and registered as Resources. These resources are utilized in all of our experiments.
Different occlusions are tested in the following test series. The experiments are executed on the AR face database, using several of the protocols that are associated with the database.
Under the hood, the ./bin/verify.py script is used to execute the algorithms. This time – as we need to run experiments with different protocols – there is a fourth step, which needs to be executed:
$ ./bin/occlusion -vv -x preprocess
$ ./bin/occlusion -vv -x train
$ ./bin/occlusion -vv -x execute
$ ./bin/occlusion -vv -x evaluate --all
Note
During the execute and evaluate action, you might specify a range of --protocols on the command line. By default, all protocols will be executed and evaluated.
Pages 2 and 4 of the multi-page PDF plot occlusion.pdf that is generated is shown in figures 6 b) and c) of the book chapter, while the other two pages show the same results, but on the development set.
Facial expressions¶
The next test checks the algorithm performances when different facial expressions are given in the images. These experiments are executed on the multipie database using protocol E (expression):
$ ./bin/expression -vv -x preprocess
$ ./bin/expression -vv -x execute
$ ./bin/expression -vv -x evaluate --all
The multi-page PDF plot expressions.pdf that is generated shows the equal error rates on the development set of Multi-PIE, and the half total error rates on the evaluation set, where the latter can be found in figure 7 c) of the book chapter.
Facial pose¶
One big problem of all face recognition algorithms is the presence of non-frontal facial poses. This series of tests try to estimate, how different poses affects the face recognition algorithms. Here, we test 13 different poses from left profile over frontal pose to right profile, using steps of 15°, using the Multi-PIE protocol P (pose):
$ ./bin/pose -vv -x preprocess
$ ./bin/pose -vv -x execute
$ ./bin/pose -vv -x evaluate --all
Note
Due to a bug in the script, the original Graphs experiments in the book chapter were executed using the wrong preprocessing, Tan-Triggs, and not with I-Norm-LBP. In the present scripts, this bug has been fixed, and the results of the Graphs algorithm might differ from the ones reported in the book chapter.
The multi-page PDF plot poses.pdf that is generated shows the equal error rates on the development set of Multi-PIE, and the half total error rates on the evaluation set, where the latter can be found in figure 7 d) of the book chapter.
Experiments on image databases¶
To evaluate the algorithms on other often-used image databases, we run several experiments on various databases. Note that, since we optimized the configurations to BANCA, the results on the other databases only serve as baselines, and optimizing the configurations to those databases will surely improve recognition accuracy.
Here, we run experiments on two image databases, MOBIO and LFW. Each of the databases has several protocols, which are all executed sequentially.
MOBIO¶
The mobio database provides two protocols male and female. The sequence:
$ ./bin/image_databases -vv -d mobio -x preprocess
$ ./bin/image_databases -vv -d mobio -x execute
will run the image-based experiments on the MOBIO database.
Note
Here, we do not evaluate the mobio database, though you could do that using ./bin/image_databases -vv -d mobio -x evaluate --all. The resulting plot is not included in the book chapter, but the mobio image experiments are evaluated together with the mobio video experiments, see below.
Since the execution of the ISV experiments on mobio (particularly the projector training) is taking a lot of time and a lot of memory, the usage of the --parallel or the --grid option is recommended. In this case, we rely on the parallel implementation of ISV by using the ./bin/verify_isv.py script.
Warning
Some of the images for the two protocols overlap. Hence, there might be a problem when using the --grid option, i.e., we have observed issues when the preprocessing of the files from both protocols is run at the same time. The --parallel option does not have this issue, it is safe to be used.
Note
The training of the MOBIO database is independent of the protocol and need to be done only once. We might have set up the scripts to take care of that, but instead we just rely on the fact that any file that has already been generated is not regenerated. Thus, although the training procedure is called twice, it will only be executed once. This might – again – not be true if the training is run in parallel.
LFW¶
The experiments of the lfw database are run on view2 only – we skip view 1 as we do not optimize the parameters to the database; which is the only purpose of view1. Still, 10 different face recognition experiments are run for each algorithm, this time requiring to re-train the algorithms for each of the 10 fold‘s.
To run all 10 experiments of the lfw database, a single sequence of call is sufficient:
$ ./bin/image_databases -vv -d lfw -x preprocess
$ ./bin/image_databases -vv -d lfw -x execute
$ ./bin/image_databases -vv -d lfw -x evaluate --all
The resulting plot databases.pdf can be found in figure 8 b) of the book chapter. The additional results of CS-KDA are not re-computed in our experiments, but they are hard-coded in the evaluation script.
Warning
The LFW database is huge. Running experiments might require a very long time and an incredible amount of memory especially since each of the folds need to re-train the algorithms. Therefore, running the experiments in --parallel or --grid mode. As before, for ISV we use the ./bin/verify_isv.py script to execute the algorithm – as otherwise the 10 ISV trainings would have taken month to finish.
Warning
Again, the images for the protocols overlap, yielding the same problem with the --grid option as for mobio.
Experiments on video databases¶
The last set of experiments utilizes video frames instead of images. Particularly, we select 1, 3, 10 and 20 frames from each video (using a simple frame selection strategy), and combine features from several frames to improve face recognition. We make use of the bob.bio.video framework, which defines simple wrapper classes that allow to use any image-based preprocessor, extractor or algorithm to work on videos.
As we do not have the required annotations in the databases, we perform a face detection and a facial landmark detection in the video frames.
Note
The number of frames that are selected can be varied with the --number-of-frames argument of the ./bin/video_databases script. Compared to the experiments on the image databases, the execution time will increase linearly with the number of frames that are selected. Also the face detector might require some additional time. Hence, we recommend to execute the experiments using a single frame (--number-of-frames 1) first, and see how long these experiments take. According to that you might estimate the time of the remaining experiments.
Note
When in the recommended --parallel or --grid mode, ISV experiments once again rely on ./bin/verify_isv.py.
MOBIO¶
For mobio, the sequence:
$ ./bin/video_databases -vv -d mobio -x preprocess -n 1
$ ./bin/video_databases -vv -d mobio -x execute -n 1
will run the experiments using a single frame (-n 1) on the MOBIO video database, on both protocols female and male in sequence. Afterward, please replace the 1 with 3, 10 and 20 in both calls.
When finished, you can evaluate the whole set of mobio experiments, i.e., the image and video experiments using:
$ ./bin/video_databases -vv -d mobio -x evaluate --all
to create the multi-page PDF file videos.pdf, which includes the equal error rates on the development set and the half total error rates on the evaluation set for all the algorithms and for both protocols female and male. The two pages that correspond to the evaluation set can be found in figure 9 b) and c) of the book chapter.
Warning
The overlap of the videos between the protocols is again valid and, thus, the experiments using the --grid option might fail.
YouTube¶
The experiments of the youtube database are split up into 10 protocols again, each of them requiring to re-train the algorithms. To run all 10 experiments of the youtube database using a single frame, call:
$ ./bin/video_databases -vv -d youtube -x preprocess -n 1
$ ./bin/video_databases -vv -d youtube -x execute -n 1
Again, repeat this for 3, 10 and 20 frames. After all the experiments have finished (or even only after parts of them have finished, you will see empty spaces in the plots), you can call:
$ ./bin/video_databases -vv -d youtube -x evaluate --all
to evaluate the experiments and generate the plot shown in figure 8 c) in the book chapter.
Algorithm run-time¶
The last set of experiments is used to measure the time that the algorithms need in specific parts of execution. Since all image pre-processing algorithms seem to need approximately the same time, which is predominated by the disk access time, we do not measure the pre-processing time.
Simply run:
$ ./bin/timimg -vv -x preprocess
$ ./bin/timimg -vv -x execute
$ ./bin/timimg -vv -x evaluate --all
to generate the latex file timing.tex that is included to constitute table 2 a) of the book chapter. This file can be imported into a LaTeX table, in a manner like:
\newcolumntype{R}{@{\quad}X<{\raggedleft}@{\quad}}
\begin{tabularx}{.9\textwidth}{p{.15\textwidth}|*{5}{|R}}
\input{timing}
\end{tabularx}
Note
As these experiments are mainly used to measure algorithm runtime, they can not be parallelized and, thus, the --grid and --parallel options are not valid for the ./bin/timing script.
Obviously, the times that you will obtain will differ from the ones that we report in the book chapter, as they depend on the machine that you are running on.
Note
The face recognition results of this execution are actually taken to create the last plot in the Configuration Optimization plot, see note above.