pymodelfit.core.FunctionModel2DScalarDeformedRadial¶
- class pymodelfit.core.FunctionModel2DScalarDeformedRadial(rmodel, atob=1, pa=0)[source]¶
Bases: pymodelfit.core.FunctionModel2DScalar
A FunctionModel2DScalar with a profile that is flattened along an axis but has a radial form if unflattened
atob is a ratio of the large to small axis, and pa is the position angle measured in radians from the x-axis to the y-axis (e.g. west to north)
Methods
__init__(rmodel[, atob, pa]) addTransform(input, output, func) Register a function as a transform from one coordinate system to another. cartesian_to_polar(x, y[, degrees]) Converts arrays in 2D rectangular Cartesian coordinates to polar chi2Data([x, y, weights, ddof]) Computes the chi-squared statistic for the data assuming this model. f(arrin, *params) fitData([x, y, fixedpars, weights, ...]) Fit the provided data using algorithms from scipy.optimize, and adjust the model parameters to match. getCov() Computes the covariance matrix for the last fitData() call. getFluxSize([flux, frac, mode, cen, v0, ...]) Compute the radius/area enclosing a specified amount of flux. getMCMC(x, y[, priors, datamodel]) Generate an Markov Chain Monte Carlo sampler for the data and model. gradient(x) integrateCartesian(xl, xu, yl, yu, **kwargs) Integrates the function in a rectangular area defined in rectangular coordinates. integrateCircular(outr[, inr, theta]) Integrates using a circular jacobian from inr to outr intrinsic_to_observed_ellipticity(ei, i[, ...]) Converts intrinsic ellipticity to observed where 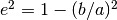
inv(output, *args, **kwargs) Compute the inverse of this model for the requested output. isVarnumModel() Determines if the model represented by this class accepts a variable number of parameters (i.e. observed_to_intrinsic_ellipticity(eo, i[, ...]) Converts observed ellipticity to intrinsic where 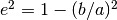
pixelize(xl, xu, yl, yu[, nx, ny, sampling]) Generates a 2D array of the model smoothed/integrated over a certain plot([datarange, nx, ny, clf, cb, data, log]) Plots the model over the range requested using the matplotlib.pyplot.imshow() function. plot3d([datarange, nx, ny, clf, cb, data]) Generate a 3D plot of the model using mayavi. plotResiduals([x, y, fmt, xaxis, clf, relative]) Plots residuals between data and the model for a variety of projections. polar_to_cartesian(r, t[, degrees]) Converts arrays in 2D polar coordinates to rectangular cartesian resampleFit([x, y, xerr, yerr, bootstrap, ...]) Estimates errors via resampling. residuals([x, y, retdata]) Compute residuals of the provided data against the model. stdData([x, y]) Determines the standard deviation of the model from data. transformCoordinates(x[, incoordsys, ...]) Transform from the input coordinate system into that defined for the model function. Attributes
data The fitting data for this model. defaultparval int(x=0) -> int or long e ellipticity given by e^2 = 1-(b/a)^2 errors Error on the data. fittype str(object=’‘) -> string fittypes A Sequence of the available valid values for the fittype fixedpars tuple() -> empty tuple inc inclination angle of object in radians - maps onto atob incdeg inclination angle of object in degrees - maps onto atob incoordsys str(object=’‘) -> string padeg params A tuple of the parameter names. pardict A dictionary mapping parameter names to the associated values. parvals The values of the parameters in the same order as params rangehint weightstype Determines the statistical interpretation of the weights in data. - addTransform(input, output, func)¶
Register a function as a transform from one coordinate system to another.
Parameters: - input (string) – The name for the input system of this function.
- output (string) – The name for the output system of this function.
- func (callable) – The function to perform the transform. It should take one argument, an array with at least 2 dimensions where the first dimension is defined by the coordinate system.
- cartesian_to_polar(x, y, degrees=False)¶
Converts arrays in 2D rectangular Cartesian coordinates to polar coordinates.
Parameters: - x (numpy.ndarray) – First cartesian coordinate
- y (numpy.ndarray) – Second cartesian coordinate
- degrees (boolean) – If True, the output theta angle will be in degrees, otherwise radians.
Returns: (r,theta) where theta is measured from the +x axis increasing towards the +y axis
- chi2Data(x=None, y=None, weights=None, ddof=1)¶
Computes the chi-squared statistic for the data assuming this model.
Parameters: - x (array-like or None) – Input data value or None to use stored data
- y (array-like or None) – Output data value or None to use stored data
- weights (array-like or None) – Weights to adjust chi-squared, typically for error bars. Statistically interpreted based on the weightstype attribute. If None, any stored data will be used.
- ddof (int) – Delta Degrees of Freedom. The divisor used for the reduced chi-squared is n-m-ddof, where N is the number of points and m is the number of parameters in the model.
Returns: tuple of floats (chi2,reducedchi2,p-value)
- data None¶
The fitting data for this model. Should be either None, or a tuple(datain,dataout,weights). Note that the weights are interpreted statistically as errors based on the weightstype attribute.
- e None¶
ellipticity given by e^2 = 1-(b/a)^2
- errors None¶
Error on the data. Sets the weights on data assuming the interpretation for errors given by weightstype. If data is None/missing, a TypeError will be raised.
- fitData(x=None, y=None, fixedpars='auto', weights=None, savedata=True, updatepars=True, fitf=False, contraction='sumsq', **kwargs)¶
Fit the provided data using algorithms from scipy.optimize, and adjust the model parameters to match.
The fitting technique is sepcified by the fittype attribute of the object, which by default can be any of the optimization types in the scipy.optimize module (except for scalar minimizers)
The full fitting output is available in lastfit attribute after this method completes.
Parameters: - x (array-like) – The input values at which to evaluate the model. Valid shapes are those that this model will accept.
- y (array-like) – The expected output values for the model at the given x values. Valid shapes are those that this model will output.
- fixedpars (sequence of strings, ‘auto’ or None) – Parameter names to leave fixed. If ‘auto’ the fixed parameters are inferred from self.fixedpars (or all will be free parameters if self.fixedpars is absent). If None, all parameters will be free.
- weights –
Weights to use for fitting, statistically interpreted as inverse errors (not inverse variance). May be one of the following forms:
- None for equal weights
- an array of points that must match the output
- a 2-sequence of arrays (xierr,yierr) such that xierr matches the x-data and yierr matches the y-data
- a function called as f(params) that returns an array of weights that match one of the above two conditions
- savedata (bool) – If True, x,`y`,and weights will be saved to data. Otherwise, data will be discarded after fitting.
- updatepars (bool) – If True, sets the parameters on the object to the best-fit values.
- fitf (bool) – If True, the fit is performed directly against the f() method instead of against the model as evaluated if called (as altered using setCall()).
- contraction –
Only applies for optimize-based methods and is the technique used to convert vectors to figures of merit. this is composed of multiple string segments:
- these will be applied to each element of the vector first:
- ‘sq’: square
- ‘abs’:absolute value
- ‘’:raw value
- the contraction is performed after:
- ‘sum’: sum of all the elements
- ‘median’: median of all the elements
- ‘mean’: mean of all the elements
- ‘prod’: product of all the elements
- optionally,the string can include ‘frac’ to use the fractional version of the differnce vector instead of raw values. For the leastsq method, this is the only applicable value
- these will be applied to each element of the vector first:
kwargs are passed into the fitting function.
Returns: array of the best fit parameters Raises ModelTypeError: If the output of the model does not match the shape of y. See also
- fittypes None¶
A Sequence of the available valid values for the fittype attribute. (Read-only)
- getCov()¶
Computes the covariance matrix for the last fitData() call.
Returns: The covariance matrix with variables in the same order as params. Diagonal entries give the variance in each parameter. Warning
This is not guaranteed to work for custom fit-types, but will always work with the default (leastsq) fit.
- getFluxSize(flux=0.5, frac=True, mode='radial', cen=(0, 0), v0=1, minfunc='fmin', intkwargs=None, **kwargs)¶
Compute the radius/area enclosing a specified amount of flux.
Parameters: - flux (float) – Specifies the flux value at which to compute the size (in model units if frac is False).
- frac (bool) – If True, interprets flux as a fraction of the total flux instead of an absolute model flux value.
- mode –
Specifies the way to compute flux. Can be:
- ‘radial’ : computes the radius enclosing the specified flux - return value in this case is a single scalar
- ‘square’ : computes the square enclosing the specified flux - return value in this case is a single scalar with the box length
- ‘rectangular’ : computes the box enclosing the specified flux - return value in this case is a 2-tuple (xsize,ysize)
- cen (2-tuple (x,y)) – Specifies the center to assume. Currently this must be (0,0) for ‘radial’ profiles.
- v0 (float) – The initial guess at which to start (for methods that require it).
- minfunc (string) – The name of a function in scipy.optimize that should do the minimizing.
- intkwargs (dictionary) – Keyword arguments for the integrate method.
kwargs are passed into the minimization function
Returns: The location at which the flux enclosed is given by flux as specified by the mode argument.
- getMCMC(x, y, priors={}, datamodel=None)¶
Generate an Markov Chain Monte Carlo sampler for the data and model. This function requires the PyMC package for the MCMC internals and sampling.
Parameters: - x (array-like) – Input data value
- y (array-like) – Output data value
- priors (dictionary) –
Maps parameter names to the priors to assume for that parameter. There must be an entry for every parameter in the model. The prior specification can be in any of the following forms:
- A pymc.Stochastric object
- A 2-tuple (lower,upper) for a uniform prior
- A scalar > 0 to use a gaussian prior of the provided width centered at the current value of the parameter
- 0 for a Poisson prior with k set by the current value of the parameter
- datamodel –
Specifies the model to assume for the fitting data points. May be any of the following:
- None
- A normal distribution with sigma given by the data’s standard deviation.
- A tuple (dist,dataname,kwargs)
- The first element is the pymc.distribution to be used as the distribution representing the data and the second is the name of the argument to be associated with the FunctionModel1D’s output, and the third is kwargs for the distribution (“observed” and “data” will be ignored, as will the data argument)
- A sequence
- A normal distribution is used with sigma for each data point specified by the sequence. The length must match the model.
- A scalar
- A normal distribution with the given standard deviation.
Raises ValueError: If a prior is not provided for any parameter.
Returns: A pymc.MCMC object ready to sample for this model.
- inc None¶
inclination angle of object in radians - maps onto atob
- incdeg None¶
inclination angle of object in degrees - maps onto atob
- integrateCartesian(xl, xu, yl, yu, **kwargs)¶
Integrates the function in a rectangular area defined in rectangular coordinates.
Parameters: kwargs are passed into scipy.integrate.dblquad()
- integrateCircular(outr, inr=0, theta=(0, 6.283185307179586), **kwargs)¶
Integrates using a circular jacobian from inr to outr
Parameters: kwargs are passed into scipy.integrate.dblquad()
- static intrinsic_to_observed_ellipticity(ei, i, degrees=True)[source]¶
Converts intrinsic ellipticity to observed where
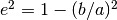
Parameters: - ei (float or array-like) – intrinsic ellipticity
- i (float or array-like) – inclination angle
- degrees – if True, the inclination is assumed to be in degrees
Returns: observed ellipticity
- inv(output, *args, **kwargs)¶
Compute the inverse of this model for the requested output.
Parameters: output – The output value of the model at which to compute the inverse. Returns: The input value at which the model produces output Raises ModelTypeError: If the model is not invertable for the provided data set.
- classmethod isVarnumModel()¶
Determines if the model represented by this class accepts a variable number of parameters (i.e. number of parameters is set when the object is created).
Returns: True if this model has a variable number of parameters.
- static observed_to_intrinsic_ellipticity(eo, i, degrees=True)[source]¶
Converts observed ellipticity to intrinsic where
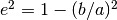
Parameters: - eo (float or array-like) – observed ellipticity
- i (float or array-like) – inclination angle
- degrees – if True, the inclination is assumed to be in degrees
Returns: intrinsic ellipticity
- params None¶
A tuple of the parameter names. (read-only)
- pardict None¶
A dictionary mapping parameter names to the associated values.
- pixelize(xl, xu, yl, yu, nx=None, ny=None, sampling=None)¶
Generates a 2D array of the model smoothed/integrated over a certain pixel size.
Parameters: - xl (float) – left edge of pixelized region
- xu (float) – right edge of pixelized region
- yl (float) – lower edge of pixelized region
- yu (float) – upper edge of pixelized region
- nx (int or None) – Number of horizontal pixels. If None, assumes xl and xu are in nearest pixel numbers.
- ny (int or None) – Number of vertical pixels. If None, assumes yl and yu are in nearest pixel numbers.
- sampling (None or int) – If None, each pixel will be computed by integrating the model over the area of the pixel. Otherwise, sampling gives the factor to multiply nx and ny by to get the total number of “virtual pixels” that will be averaged to get the actual result at the given grid point. If <=1, this is equivalent to directly sampling the model.
Returns: An nx X ny array with the pixelized model.
Note
The attribute lastintegrate stores the result of the integrations if sampling is None, with lastintegrate[0] storing the result and lasteintegrate[1] storing the error on the integral for each pixel.
- plot(datarange=None, nx=100, ny=100, clf=True, cb=True, data='auto', log=False, **kwargs)¶
Plots the model over the range requested using the matplotlib.pyplot.imshow() function.
Parameters: - datarange (4-tuple or None) – Specifies the range to plot in the form (xl,xu,yl,yu). If None, it will be inferred from the data.
- nx (int) – determines the number of pixels in the x-direction
- ny (int) – determines the number of pixels in the y-direction
- clf (bool) – If True, the figure will be cleared before plotting.
- data (array-like, None, or ‘auto’) – Data to be plotted along with the model. If None, no data will be plotted, or if ‘auto’, it will be taken from data (if present).
- log (bool or string) – Can be False, True (natural log), ‘10’ (base-10), ‘mag’, or ‘mag##’ (pogson magnitudes with ## as zeropoint)
kwargs are passed into matplotlib.pyplot.imshow()
Raises ValueError: if data is not present and datarange is None
- plot3d(datarange=None, nx=100, ny=100, clf=True, cb=True, data='auto', **kwargs)¶
Generate a 3D plot of the model using mayavi.
See plot() for meaning of the arguments
- plotResiduals(x=None, y=None, fmt='or', xaxis='r', clf=True, relative=False)¶
Plots residuals between data and the model for a variety of projections. If x and y are None, the residuals will be inferred from the last-fit data. For details on how residuals are computed see FunctionModel.residuals()
Parameters: - x (array-like) – input data
- y (array-like) – output data
- fmt – fmt argmument for matplotlib.pyplot.ploy()
- xaxis (string) – Sets the plane in which to plot residuals. Can be ‘r’,’theta’,’x’, or ‘y’.
- clf (bool) – If True, the plot will be cleared before plotting.
- relative – If true, the plot will be relative residuals (e.g. residual/model) instead of the absolute residuals.
- polar_to_cartesian(r, t, degrees=False)¶
Converts arrays in 2D polar coordinates to rectangular cartesian coordinates.
Note that the spherical coordinates are in physicist convention such that (1,0,pi/2) is x-axis.
Parameters: - r (numpy.ndarray) – Radial coordinate
- t (numpy.ndarray) – Azimuthal angle from +x-axis increasing towards +y-axis
- degrees (boolean) – If True, the input angles will be in degrees, otherwise radians.
Returns: arrays (x,y)
- resampleFit(x=None, y=None, xerr=None, yerr=None, bootstrap=False, modely=False, n=250, prefit=True, medianpars=False, plothist=False, **kwargs)¶
Estimates errors via resampling. Uses the fitData function to fit the function many times while either using the “bootstrap” technique (resampling w/replacement), monte carlo estimates for the error, or both to estimate the error in the fit.
Parameters: - x (array-like or None) – The input data - if None, will be taken from the data attribute.
- y (array-like or None) – The output data - if None, will be taken from the data attribute.
- xerr (array-like, callable, or None) – Errors for the input data (assumed to be normally distributed), or if None, will be taken from the data. Alternatively, it can be a function that accepts the input data as the first argument and returns corresponding monte carlo sampled values.
- yerr (array-like, callable, or None) – Errors for the output data (assumed to be normally distributed), or if None, will be taken from the data. Alternatively, it can be a function that accepts the output data as the first argument and returns corresponding monte carlo sampled values.
- bootstrap (bool) – If True, the data is also resampled (with replacement).
- modely – If True, the fitting data will be generated by offseting from model values (evaluated at the x-values) instead of y-values.
- n (int) – The number of times to draw samples.
- prefit (bool) – If True, the data will be fit without resampling once before the samples are recorded.
- medianpars (bool) – If True, the median from the histogram will be set as the value for the parameter.
- plothist (bool or string) – If True, histograms will be plotted using matplotlib.pyplot.hist() for each of the parameters, or if it is a string, only the histogram for the requested parameter will be shown.
kwargs are passed into fitData
Returns: (histd,cov) where histd is a dictionary mapping parameters to their histograms and cov is the covariance matrix of the parameters in parameter order. Note
If x, y, xerr, or yerr are provided, they do not overwrite the stored data, unlike most other methods for this class.
- residuals(x=None, y=None, retdata=False)¶
Compute residuals of the provided data against the model. E.g.
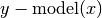 .
.Parameters: Returns: Residuals of model from y or if retdata is True, a tuple (x,y,residuals).
Return type: array-like
- stdData(x=None, y=None)¶
Determines the standard deviation of the model from data. Data can either be provided or (by default) will be taken from the stored data.
Parameters: Returns: standard deviation of model from y
- transformCoordinates(x, incoordsys=None, outcoordsys=None)¶
Transform from the input coordinate system into that defined for the model function.
Parameters: - incoordsys (string or None) – The input coordinate system name to use for this model. If None, incoordsys will be used.
- outcoordsys (string) – The output coordinate system name to use for this model. If None, the standard for this model will be used.
if incoordsys is None, self.`incoordsys will be used
- weightstype None¶
Determines the statistical interpretation of the weights in data. Can be:
- ‘ierror’
Weights act as inverse errors (default)
- ‘ivar’
Weights act as inverse variance
- ‘error’
Weights act as errors (non-standard - this makes points with larger error bars count more towards the fit).
- ‘var’
Weights act as variance (non-standard - this makes points with larger error bars count more towards the fit).